

The University of Washington Nanopore Sequencing Core provides DNA and RNA sequencing services on the Oxford Nanopore Technologies platform. For collaborators, we perform library preparation and sequencing from a variety of sample inputs, including blood, saliva, buccal swabs, and cell pellets. We also accept direct DNA or RNA submissions.
This page provides detailed information on available services and how to prepare, package, and ship your samples. Before submitting samples, we ask that you contact us for a consultation.
Our Sample Submission Process:
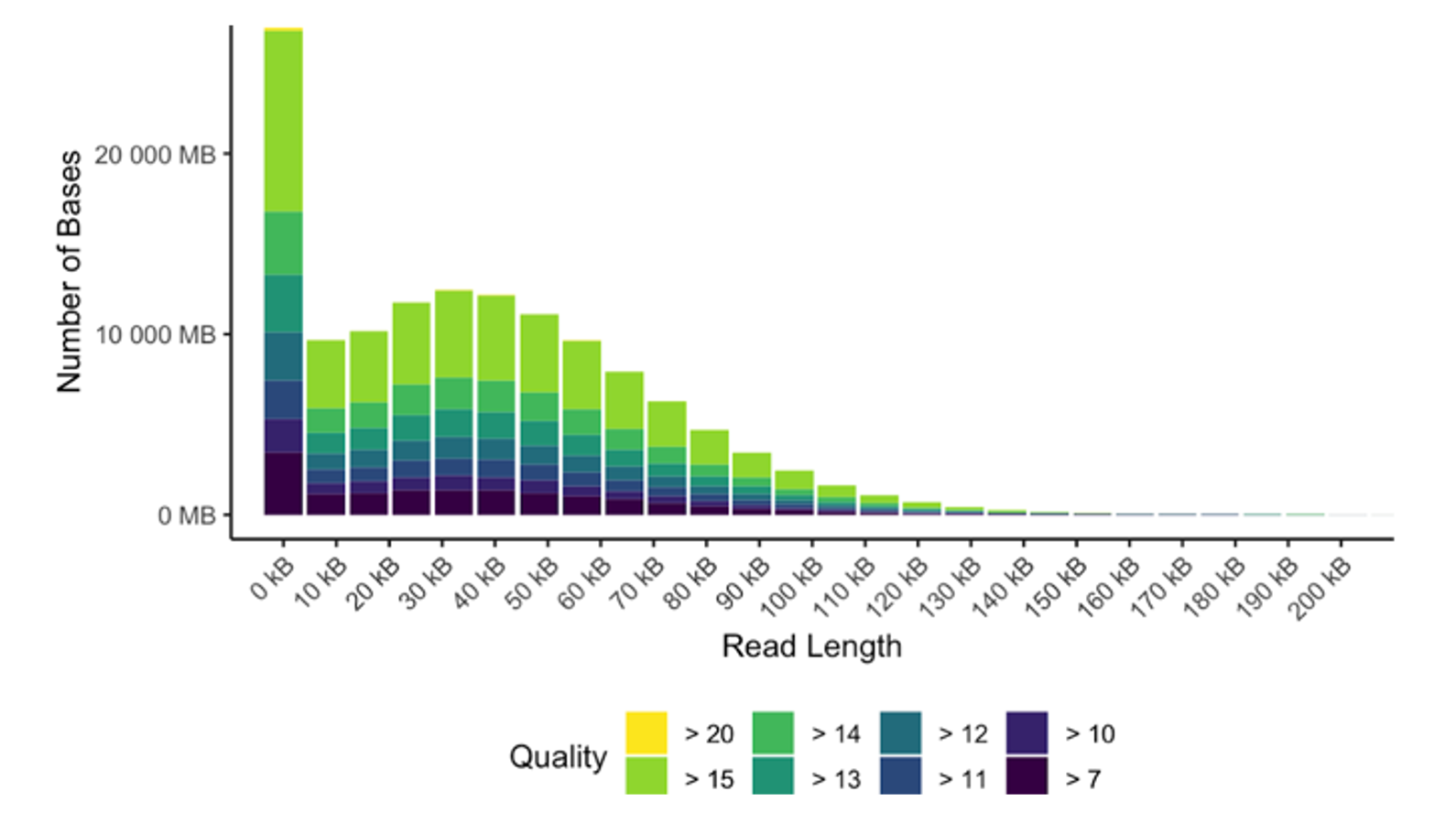
Nanopore sequencing is a third-generation sequencing technology that allows for analysis of DNA and RNA sequences without amplification or modification. As nucleic acids pass through protein nanopores, changes in electric current are detected and subsequently decoded into bases, preserving methylation and modifications to the original sequences. With the PromethION instrument, we generate high-quality datasets with average read lengths of 10–60 kb — suitable for long-read applications such as de novo assembly, structural variant discovery, splice variant discovery, and methylation analysis.
We offer cell culture services for projects requiring fibroblast or lymphoblastoid cell lines. We can culture cells from patient-derived samples, ensuring high-quality growth and expansion for downstream genetic analysis or sequencing. Our team can provide guidance on cell line creation and maintenance, helping ensure that your project progresses smoothly.
DNA and RNA extraction and purification services are available on material inputs including cell pellets, saliva, buccal swabs, and blood. Please see Sample Preparation below for more detail. Contact us to discuss tissue extractions.
For all samples, DNA will be quantified using the Qubit, a NanoDrop will be used to assess possible extraction carryover and RNA content (260:230 and 260:280 ratios, respectively), and the Agilent Femto Pulse system is used to evaluate DNA for quality and fragment length. If necessary based on QC results, sample cleanup, shearing, and/or size selection may be performed at additional cost.
Library preparation methods depend on the type of samples being sequenced. We typically use a ligation library prep for DNA sequencing and a cDNA prep for RNA sequencing, however we also offer direct RNA and rapid library preps.
We also accept already-prepared libraries for sequencing, but please be aware that we are unable to perform our typical QC on those and thus cannot guarantee their performance. Results can and do vary based on the quality of the extraction and library preparation methods used. For example, DNA sequencing of samples with protein or RNA carryover may not sequence well.
Using either a rapid or ligation library preparation, we can barcode, pool, and sequence up to 24 samples on a PromethION flow cell. Keep in mind that increasing the number of samples per flow cell will decrease the coverage of each sample. A minimum of 4 samples is required for barcoded runs.
Sequenced libraries may produce datasets of more than 120 Gb, or ~30–40x coverage of the human genome. The raw signal output for all samples will be basecalled using Dorado. We offer several sequencing strategies on the PromethION platform, including:
One hour of analysis is charged per flow cell. Samples are basecalled and run through our standard in-house analysis pipeline, which calls 5mCG/5hmCG modifications when sequencing native DNA. The calls are provided as tags in the unaligned bam files that are returned when sequencing is complete. Our standard pipeline is optimized for human samples; data for other organisms may be delivered in other formats.
For an added fee, we also offer a range of additional analysis services that can be tailored to meet the specific needs of your study. For example, our team can assist with tasks such as calling additional methylation models, filtering and prioritizing structural variants, identifying repeat expansions, analyzing other complex variants, and data interpretation support for researchers looking to analyze their own data.
We can help you determine which sequencing and analysis services are right for your project. Contact us to set up a consultation and to recieve an estimate.
| Service | Internal UW* | External Academic or Nonprofit | Industry |
|---|---|---|---|
| Cell Culture | $116.72 | $134.93 | $148.42 |
| Nucleotide Extraction | $119.62 | $138.29 | $152.11 |
| Sample QC | $55.79 | $64.49 | $70.94 |
| Sample Cleanup | $41.99 | $48.54 | $53.40 |
| Shearing | $48.85 | $56.47 | $62.11 |
| Size Selection | $99.38 | $114.88 | $162.37 |
| Library Preparation | $259.72 | $300.24 | $330.27 |
| Flow Cell | $1,238.95 | $1,432.22 | $1,575.45 |
| Barcoding – Rapid Prep (per sample) | $41.77 | $48.29 | $53.12 |
| Barcoding – Ligation Prep (per sample) | $84.16 | $97.28 | $107.01 |
| Barcoding – Pool (4–24 samples per pool) | $106.24 | $122.81 | $135.09 |
| Analysis (per hour) | $218.36 | $252.42 | $277.66 |
*Staff and faculty at the University of Washington paying with a UW Worktag receive the Internal rate; all others are charged the External or Industry rate. Rates are subject to change without notice.
Genome coverage and read length distributions in your aligned data are highly dependent on the quality of input material. The following guidelines will help ensure you receive the highest quality data possible.
We currently accept cell pellets, saliva samples, buccal swabs, and blood sample inputs, as well as DNA and RNA submissions. For other sample types, please inquire directly. Follow the guidelines below for your specific sample type.
Please submit between 1 x 10e5 and 1 x 10e7 cells per pellet. Cells must be frozen as pellets in conical tubes. Microcentrifuge tubes should be double-packed into larger conical tubes and shipped in a polystyrene box filled at least halfway with dry ice. Send these samples using next-day delivery service, for delivery between 10:00 am – 4:00 pm.*
Saliva and buccal swabs collected using DNA Genotek kits can be shipped at room temperature in padded envelopes. Our current extraction protocol uses PrepIT.L2P.
Our recommendation for stabilizing blood samples is to use a PAXgene blood DNA tube. PAXgene tubes can be shipped to us at room temperature within one week of collection. They can be shipped frozen within 2 months of collection. Frozen PAXgene tubes should be shipped horizontally.
If PAXgene tubes are not available, blood must be stabilized with EDTA, citrate, or heparin in a vacutainer. If possible, blood should not be frozen. If blood must be frozen and shipped, it should be flash frozen quickly after collection and shipped on dry ice.
Please inquire directly about pick up or drop off of fresh samples in the Seattle area.
Our current extraction protocol uses the Monarch® HMW DNA Extraction Kit for Cells & Blood for frozen blood with 500 µL to 2 mL input or the Puregene Blood Kit for fresh blood with 2+ mL of input.
For DNA submission, please run a standard or pulsed-field gel, BioAnalyzer, or TapeStation to visualize DNA quality and length before submitting. If your sample looks degraded, re-extract the sample or plan for shorter read lengths. While ~30–40x coverage of the human genome may be achieved with one PromethION flow cell, poor quality DNA will not yield these results. If submitting old, highly fragmented, or otherwise degraded DNA, you should expect ~10–20x coverage.
Additional guidelines:
Ship DNA in LoBind microcentrifuge tubes or LoBind PCR plates (Eppendorf). Pad and insulate your samples to minimize shearing during shipping.
For direct RNA submission, please contact us to discuss the guidelines.
We’re happy to help! Just send us an email.
If you would like to drop off your samples in person, we accept delivery 9 am – 11 am on weekdays.
For shipping, please plan for weekday delivery only, 10:00 am – 4:00 pm.* All samples should be delivered to:
University of Washington Nanopore Sequencing Core
c/o Joy Goffena
1705 NE Pacific St.
HSB RM H-458
Seattle, WA 98195
For all deliveries:
* Please do not send packages for first morning delivery. FedEx and UPS do not have early morning access to our building, so deliveries arriving prior to 9 am, including FedEx First Overnight deliveries, may be delayed by a day or more.
* Please do not send packages to arrive on weekends or after 4:00 pm on weekdays.
* Please do not ship samples without contacting us first.